 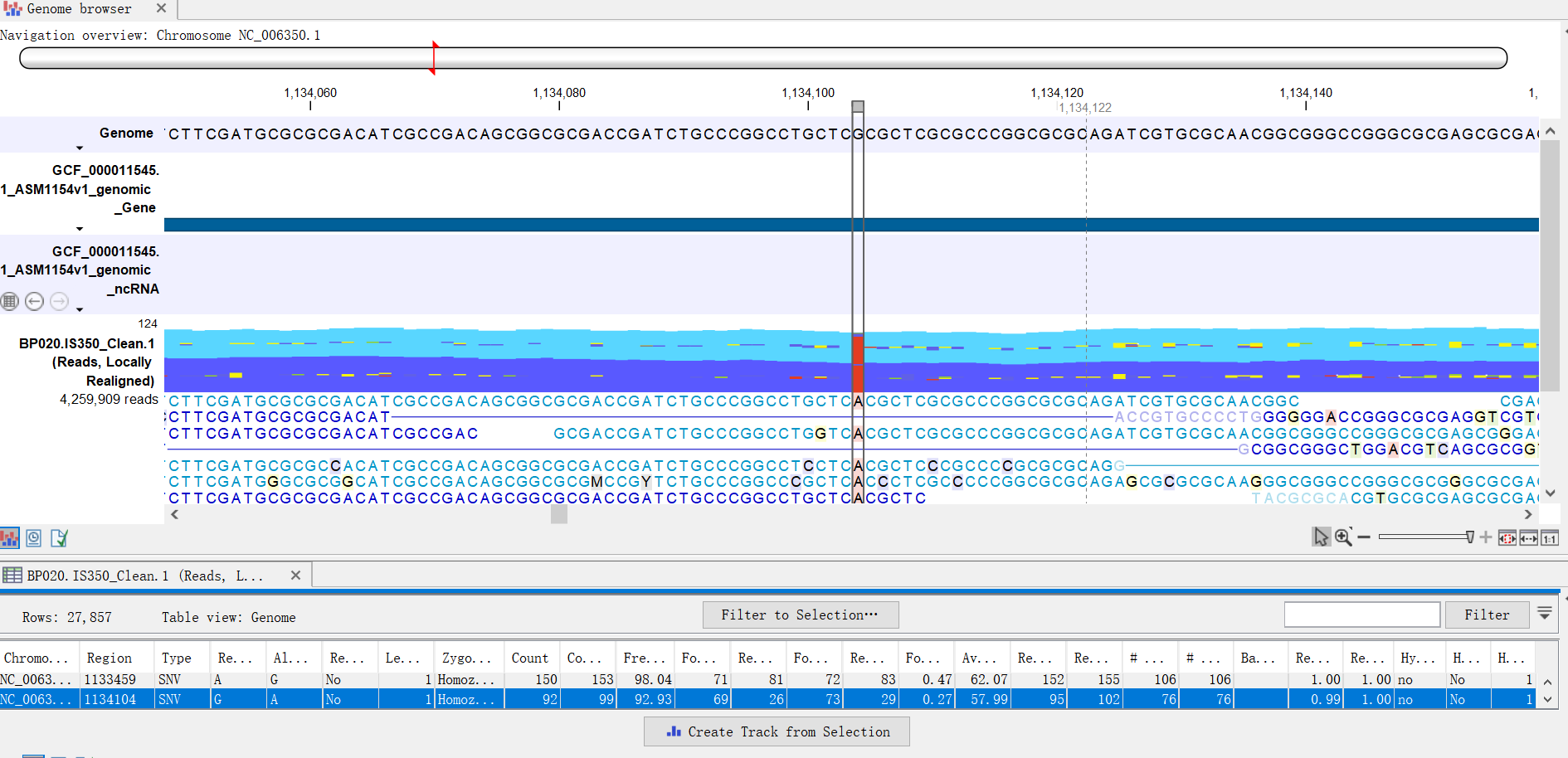
The names of the references are not checked when determining if different track objects are compatible with one another - for example, if their contents can be compared or if they be added to the same track list.Thus, if you are working with a track-based read mapping, you can just use that alongside your original track-based reference genome sequence, for example, in a Track List.Įxtra notes on working with tracks within the Workbench: The genome sequence information in the original track set is the same as that in the stand-alone Sequence List you created. You do not need to convert your reference set back to track format if you started with a set of references in track format. More information about this is provided in the link below: (You can choose to create a track-based or stand-alone read mapping during this import). Import the SAM/BAM file using the reference set with the names that match those used in the SAM/BAM file.In the image below, we change the names from R to chrR (e.g. Use the Rename Sequences in Lists functionality ( ) from the Utility Tools folder in the Toolbox to change the names appropriately.If working with more than a few sequences:.Just right click on each sequence name in turn and choose the option Rename Sequence. If working with only a few sequences, you can change the names directly in the Sequence List.Instructions on how to do this is provided in the link below: Convert the reference genome sequence track to a stand-alone sequence list if the references are initially in track format.If you have a set of reference sequences in the Workbench that use one naming sequence and your SAM/BAM file contains references using a different naming scheme, then the method below can be used to create a reference set that can be used for importing the mapping data. For example, in the case of the human genome, chromosomes in different public resources have different naming patterns, such as "chrR", "R" and "NC_00000R", where R is some integer number or a letter. The issue of reference names commonly arises when using data from resources where different naming schemes are applied. If the reference names in a SAM/BAM file do not match the reference names in the Workbench, then the easiest route is usually to change names of the reference sequences in the Workbench to match those in your SAM/BAM file. The reference sequences in the SAM/BAM file and in the Workbench must match in both name and lengths in order to be able to import mapped data. To import mapping data from a SAM or BAM file you need to already have the reference sequences in the Workbench. This FAQ covers guidelines on how to import SAM/BAM file whose reference names are different than those present in the Workbench I would be able to speak on the phone if that makes it easier to assist me, and I appreciate and help and advice you can provide.How can I import mappings from a SAM/BAM file where the reference names are different to those in the Workbench? Which tool should I use for the trimming? Once it is trimmed can I go straight to mapping, and should I use tophat or bowtie2? My question is how should I combine the data and process it in such a way that it is ready to be mapped?įrom what I've read I will change the data type to fastqsanger, concatenate the technical replicates for the forward and reverse reads and then trim the files. 
I believe that the files are in fastqsanger format. I therefore have 16 files per experimental group (forward and reverse for each of the 4 biological replicates, each run in two lanes. Each sample was also run in two separate lanes (we were told these technical replicates willl give us more information on isoform expression). 
We have 4 biological replicates from 4 experimental groups. They prepared the libraries and performed paired end sequencing. I just got back RNA seq data produced on the HiSeq 4000. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
